Engineering polyspecific VHHs with picomolar binding to SARS-CoV-2 wild type and Omicron
Scientists at Cradle optimized a VHH for simultaneous binding to SARS-CoV-2 wild type and Omicron, thermostability, and expression across three rounds. The best candidate achieved a K_D of 186 pM against wild type, 11.4 nM against Omicron, a melting temperature of 70.9 °C, and 1.88× improved expression.

Daniel

Daniel
Modality | VHH (nanobody) |
Target | SARS-CoV-2 (wild type + Omicron variant) |
Properties optimized | Binding (2×), thermostability, expression |
Rounds | 3 |
Candidates per round | 96 |
Key result | K_D 186 pM (WT), K_D 11.4 nM (Omicron), T_m 70.9 °C, 1.88× expression |
Partner | Internal |
Data availability | Detailed in CRADLE-1 whitepaper |
Context
SARS-CoV-2 variant evolution necessitates broadly neutralizing antibodies capable of binding multiple variants. Polyspecific binders are particularly valuable as the virus continues to mutate, but simultaneous optimization across multiple binding targets while maintaining biophysical properties represents a challenging multi-dimensional optimization problem.
Challenge
The template sequence, derived from phage display NGS data using Cradle's hit identification module, already exhibited reasonably good properties: K_D of 2.01 nM against wild type, K_D of 34.6 nM against Omicron, and a melting temperature of 58.5 °C. This made the optimization particularly challenging—the template was already functional, meaning most mutations would be deleterious to one or more properties, with relatively few pathways for simultaneous improvement.
The requirement for polyspecific binding creates inherent trade-offs: mutations that improve binding to wild type may reduce binding to Omicron, and vice versa. Additionally, stability and expression constraints further limit the viable sequence space.

Meet the Cradle wet lab
Discover why we, unlike other AI companies, run our own wet lab in Amsterdam to build better models.
Approach
Our scientists performed three rounds of Cradle optimization with 96 candidates each. Round 1 operated in zero-shot mode using only evolutionary context. Round 2 prioritized thermostability as the primary objective while maintaining binding and expression constraints. Round 3 shifted the primary objective to Omicron binding while enforcing a minimum T_m threshold of 65 °C.
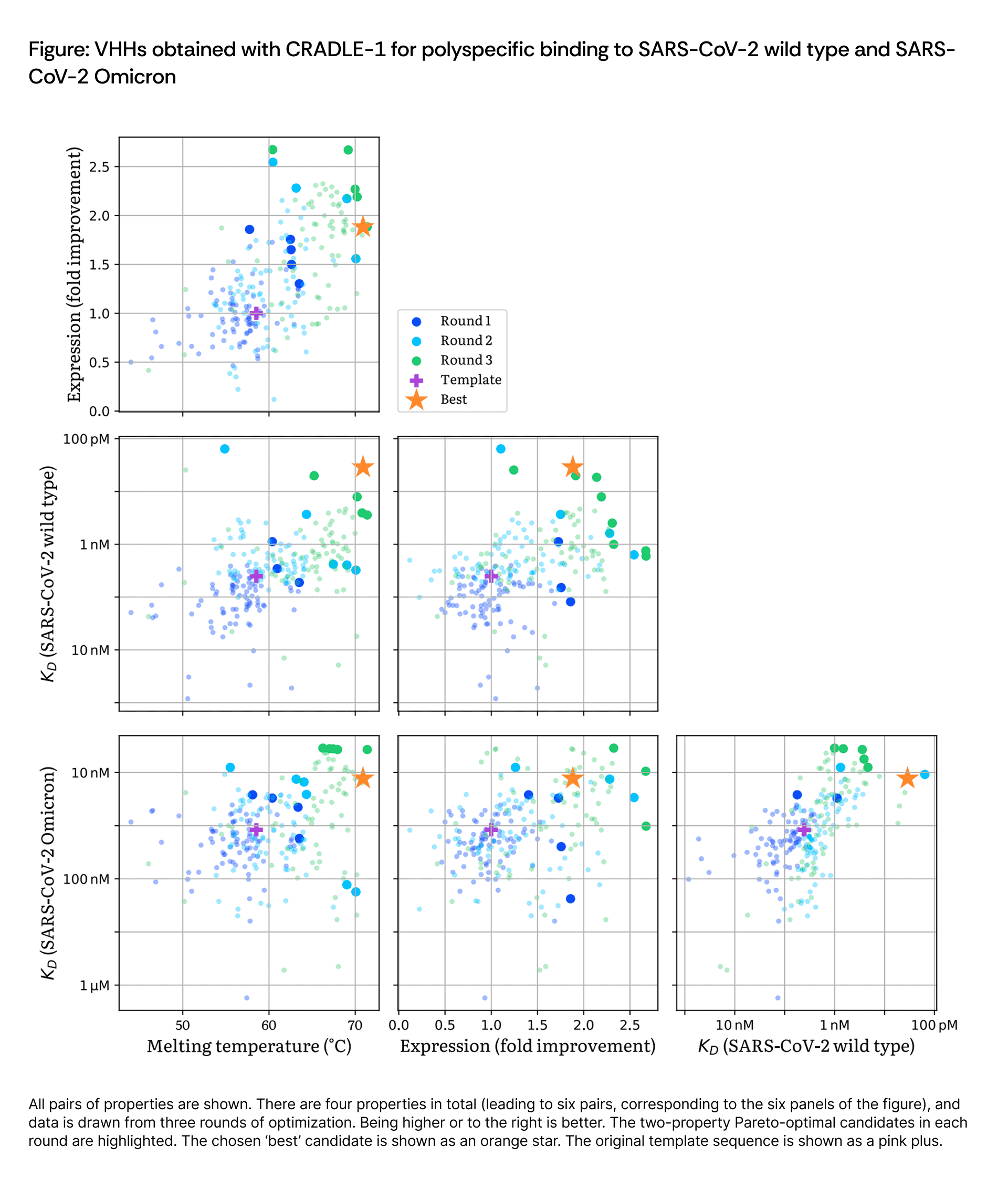
This multi-round strategy enabled active Pareto frontier exploration, strategically shifting the population toward optimal quadrants through successive iterations. The final candidate was manually selected from the Round 3 Pareto frontier, representing a reasonable trade-off across all four properties.
Results
Cradle progressively expanded the Pareto frontier across all three rounds. The final best candidate represents substantial improvements across all optimized properties:
Property | Template | Best Cradle candidate | Fold improvement |
K_D (wild type) | 2.01 nM | 186 pM | 10.8× |
K_D (Omicron) | 34.6 nM | 11.4 nM | 3.0× |
T_m | 58.5 °C | 70.9 °C | +12.4 °C |
Expression | 1× | 1.88× | 1.88× |
The Round 1 zero-shot diversification efficiently explored the functional landscape despite having no sequence-function data. Round 2 identified several variants with stability gains exceeding 10 °C over wild type, though these exhibited only moderate Omicron affinity (double-digit nM). Round 3's strategic shift to prioritize Omicron binding while maintaining thermal stability successfully delivered candidates achieving single-digit nM affinity against both targets with T_m above 70 °C.
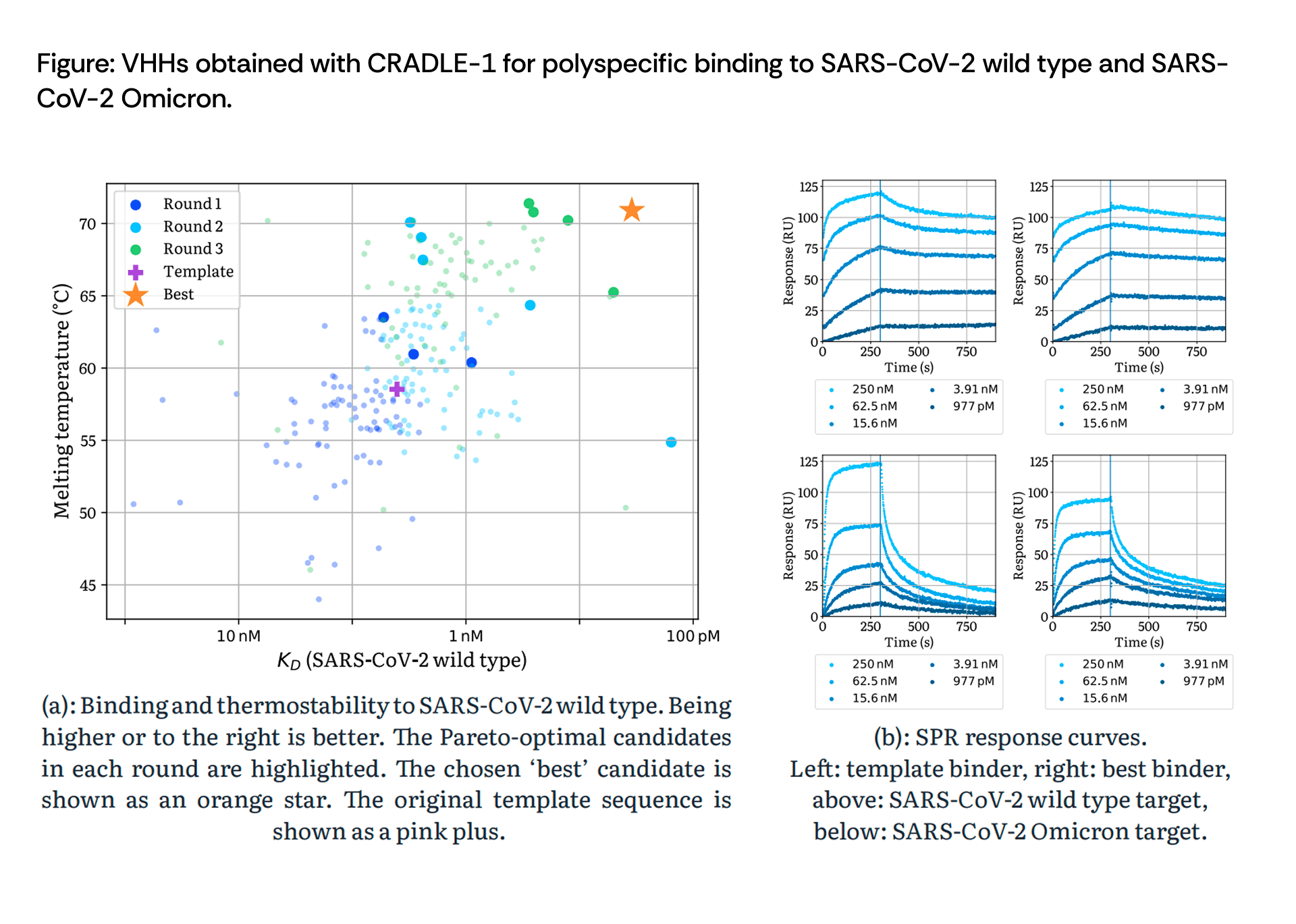
SPR response curves confirmed tight binding kinetics, with the best candidate exhibiting picomolar affinity to wild type—a 10.8-fold improvement from an already-functional starting point.
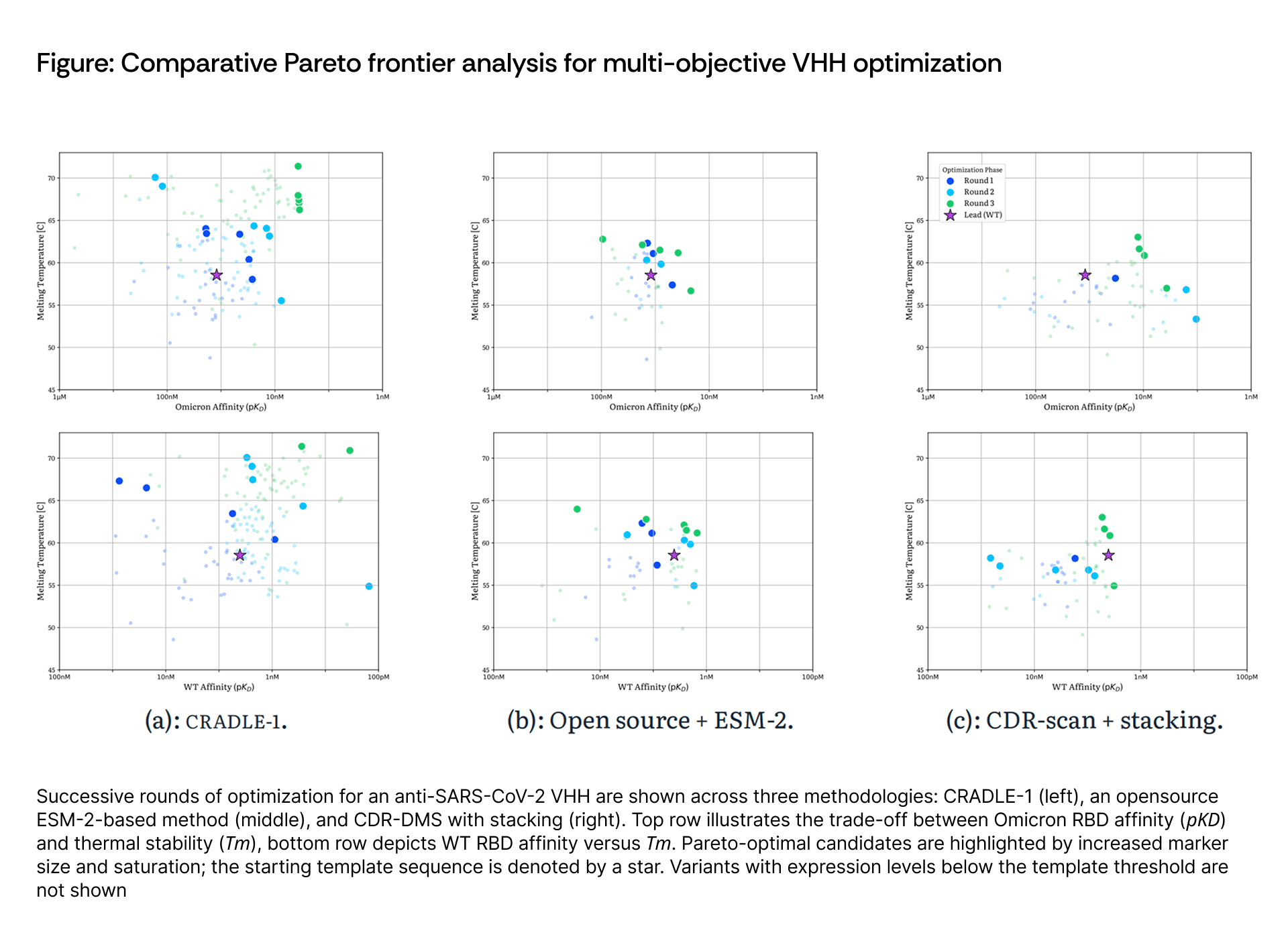
We benchmarked Cradle against two baselines: CDR3 scanning followed by recombinant stacking (classical protein engineering) and ProteusAI with ESM-2 (computational baseline). In the benchmarking phase, the ESM-2-based ProteusAI pipeline struggled with developability; 76% of its generated sequences were lost due to poor expression, whereas Cradle was the only method to demonstrate controllable, directional expansion of the Pareto frontier across successive rounds. Both baseline approaches produced largely stochastic distributions with few high-performing variants, suggesting their successes resulted from chance sampling rather than systematic landscape exploration.
What this means
This campaign demonstrates Cradle's capability for multi-property optimization of proteins that already exhibit good baseline function. Starting from a nanomolar binder and pushing to picomolar affinity while simultaneously improving three other properties illustrates the system's ability to navigate complex fitness landscapes with competing objectives. The successful zero-shot first round and progressive Pareto frontier expansion across subsequent rounds suggest that protein language models capture evolutionarily conserved binding principles that generalize across variant epitopes.
The comparison against baselines validates that Cradle's improvements stem from systematic optimization rather than stochastic search, a critical distinction for commercial lead optimization workflows.
Methods note
Cradle employed a pre-trained ESM-2 protein language model, fine-tuned via evotuning on the template's evolutionary neighborhood and via group-DPO on wet lab data. Supervised predictors trained on Round 1 and Round 2 data guided Round 2 and Round 3 generation, respectively. Binding kinetics were measured via SPR on a Carterra LSA instrument, thermostability via differential scanning fluorimetry, and expression via LabChip protein characterization. Full methodological details and baseline comparisons are available in the CRADLE-1 whitepaper.
Modality | VHH (nanobody) |
Target | SARS-CoV-2 (wild type + Omicron variant) |
Properties optimized | Binding (2×), thermostability, expression |
Rounds | 3 |
Candidates per round | 96 |
Key result | K_D 186 pM (WT), K_D 11.4 nM (Omicron), T_m 70.9 °C, 1.88× expression |
Partner | Internal |
Data availability | Detailed in CRADLE-1 whitepaper |
Context
SARS-CoV-2 variant evolution necessitates broadly neutralizing antibodies capable of binding multiple variants. Polyspecific binders are particularly valuable as the virus continues to mutate, but simultaneous optimization across multiple binding targets while maintaining biophysical properties represents a challenging multi-dimensional optimization problem.
Challenge
The template sequence, derived from phage display NGS data using Cradle's hit identification module, already exhibited reasonably good properties: K_D of 2.01 nM against wild type, K_D of 34.6 nM against Omicron, and a melting temperature of 58.5 °C. This made the optimization particularly challenging—the template was already functional, meaning most mutations would be deleterious to one or more properties, with relatively few pathways for simultaneous improvement.
The requirement for polyspecific binding creates inherent trade-offs: mutations that improve binding to wild type may reduce binding to Omicron, and vice versa. Additionally, stability and expression constraints further limit the viable sequence space.

Meet the Cradle wet lab
Discover why we, unlike other AI companies, run our own wet lab in Amsterdam to build better models.
Approach
Our scientists performed three rounds of Cradle optimization with 96 candidates each. Round 1 operated in zero-shot mode using only evolutionary context. Round 2 prioritized thermostability as the primary objective while maintaining binding and expression constraints. Round 3 shifted the primary objective to Omicron binding while enforcing a minimum T_m threshold of 65 °C.
This multi-round strategy enabled active Pareto frontier exploration, strategically shifting the population toward optimal quadrants through successive iterations. The final candidate was manually selected from the Round 3 Pareto frontier, representing a reasonable trade-off across all four properties.
Results
Cradle progressively expanded the Pareto frontier across all three rounds. The final best candidate represents substantial improvements across all optimized properties:
Property | Template | Best Cradle candidate | Fold improvement |
K_D (wild type) | 2.01 nM | 186 pM | 10.8× |
K_D (Omicron) | 34.6 nM | 11.4 nM | 3.0× |
T_m | 58.5 °C | 70.9 °C | +12.4 °C |
Expression | 1× | 1.88× | 1.88× |
The Round 1 zero-shot diversification efficiently explored the functional landscape despite having no sequence-function data. Round 2 identified several variants with stability gains exceeding 10 °C over wild type, though these exhibited only moderate Omicron affinity (double-digit nM). Round 3's strategic shift to prioritize Omicron binding while maintaining thermal stability successfully delivered candidates achieving single-digit nM affinity against both targets with T_m above 70 °C.
SPR response curves confirmed tight binding kinetics, with the best candidate exhibiting picomolar affinity to wild type—a 10.8-fold improvement from an already-functional starting point.
We benchmarked Cradle against two baselines: CDR3 scanning followed by recombinant stacking (classical protein engineering) and ProteusAI with ESM-2 (computational baseline). In the benchmarking phase, the ESM-2-based ProteusAI pipeline struggled with developability; 76% of its generated sequences were lost due to poor expression, whereas Cradle was the only method to demonstrate controllable, directional expansion of the Pareto frontier across successive rounds. Both baseline approaches produced largely stochastic distributions with few high-performing variants, suggesting their successes resulted from chance sampling rather than systematic landscape exploration.



What this means
This campaign demonstrates Cradle's capability for multi-property optimization of proteins that already exhibit good baseline function. Starting from a nanomolar binder and pushing to picomolar affinity while simultaneously improving three other properties illustrates the system's ability to navigate complex fitness landscapes with competing objectives. The successful zero-shot first round and progressive Pareto frontier expansion across subsequent rounds suggest that protein language models capture evolutionarily conserved binding principles that generalize across variant epitopes.
The comparison against baselines validates that Cradle's improvements stem from systematic optimization rather than stochastic search, a critical distinction for commercial lead optimization workflows.
Methods note
Cradle employed a pre-trained ESM-2 protein language model, fine-tuned via evotuning on the template's evolutionary neighborhood and via group-DPO on wet lab data. Supervised predictors trained on Round 1 and Round 2 data guided Round 2 and Round 3 generation, respectively. Binding kinetics were measured via SPR on a Carterra LSA instrument, thermostability via differential scanning fluorimetry, and expression via LabChip protein characterization. Full methodological details and baseline comparisons are available in the CRADLE-1 whitepaper.
Recent posts
Subscribe and get new posts and updates from Cradle straight to your inbox.
Follow Cradle
Built with ❤️ in Amsterdam & Zurich
Follow Cradle
Built with ❤️ in Amsterdam & Zurich
Follow Cradle