Using AI to produce vaccine adjuvants without cutting down rainforest
Using AI to produce vaccine adjuvants without cutting down rainforest
Every year, 10,000 Chilean soap bark trees are harvested to extract QS-21 for vaccines against malaria, shingles, and cancer. Now UC Berkeley's Keasling Lab is using AI to re-engineer plant enzymes that struggle in fermentation tanks.

Elise

Elise

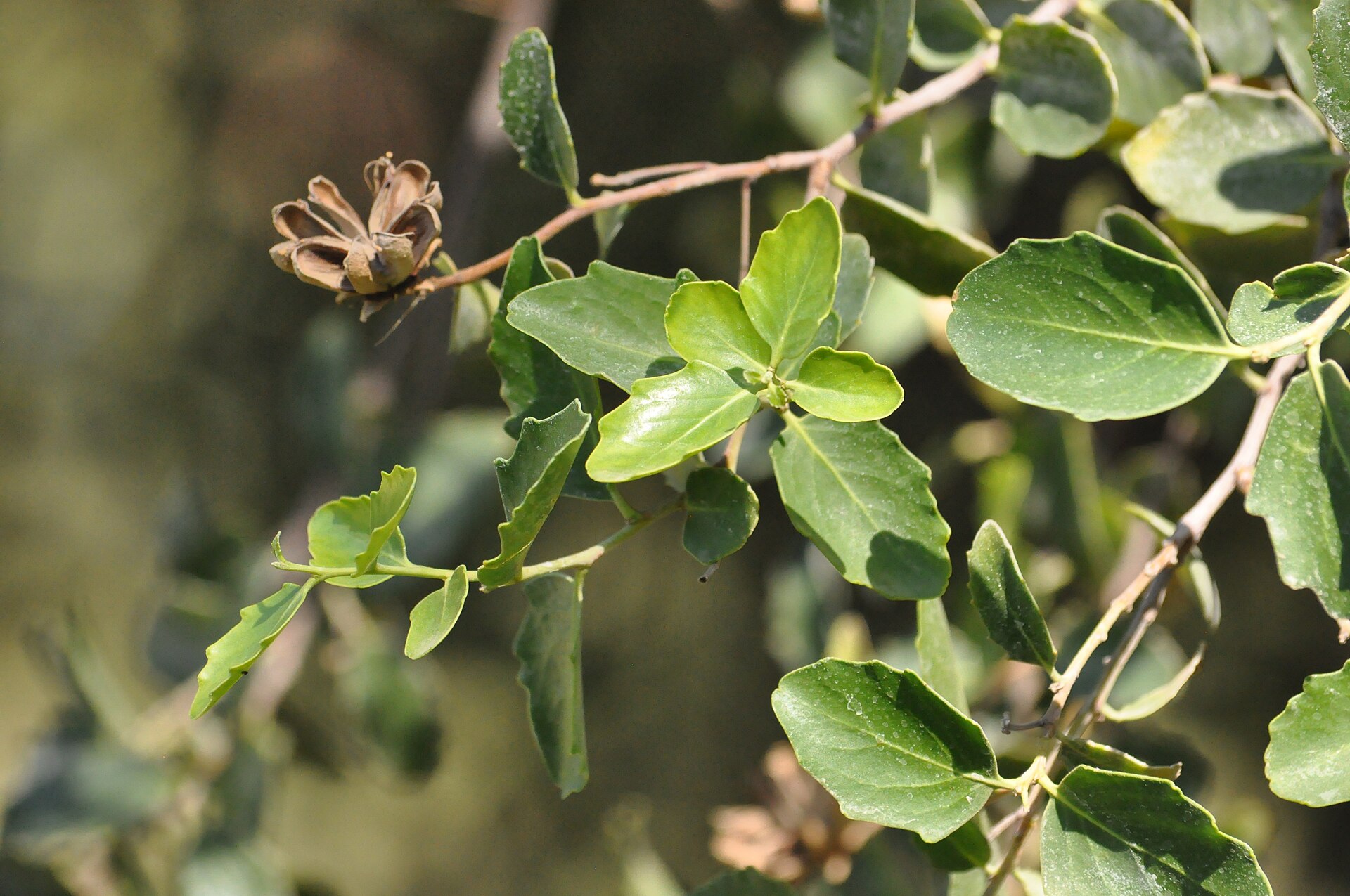
The “soap bark tree” Quillaja saponaria gets its name from the saponins in its bark—natural compounds that create a soapy lather in water. Indigenous people in the Andes have used this bark for generations as a natural remedy. Scientists later discovered that one of these saponins, a molecule called QS-21, could act as a ‘vaccine adjuvant’ to boost the immune system's response to vaccines. Today, QS-21 is a key ingredient in vaccines against shingles, malaria, and RSV.
As vaccine demand grows, so does pressure on Chile's soap bark tree populations. The trees are already threatened by deforestation and wildfires, and they grow slowly—taking decades to reach maturity. While sustainable harvesting methods have been introduced, meeting pharmaceutical demand at scale remains difficult. Biologists at UC Berkeley are working on a solution.

Maria Astolfi, born and raised in the Amazonian rainforest, is a synthetic biologist in the Keasling Lab at UC Berkeley. She’s part of a research project to replace this extraction-based supply chain with precision fermentation, and uses Cradle to accelerate their research.
Plant enzymes in a yeast cell
In precision fermentation, researchers engineer a microbe like baker’s yeast so that it becomes able to ‘brew’ a specific molecule of interest in a fermentation vat. In 2024, researchers from the Keasling lab demonstrated that it was possible to produce QS-21 using this approach (published in Nature in 2024, source). They added a copy of 38 plant genes from Quillaja saponaria into baker’s yeast, engineering its metabolism to now be able to convert renewable resources like simple sugars into this sought-after vaccine adjuvant.
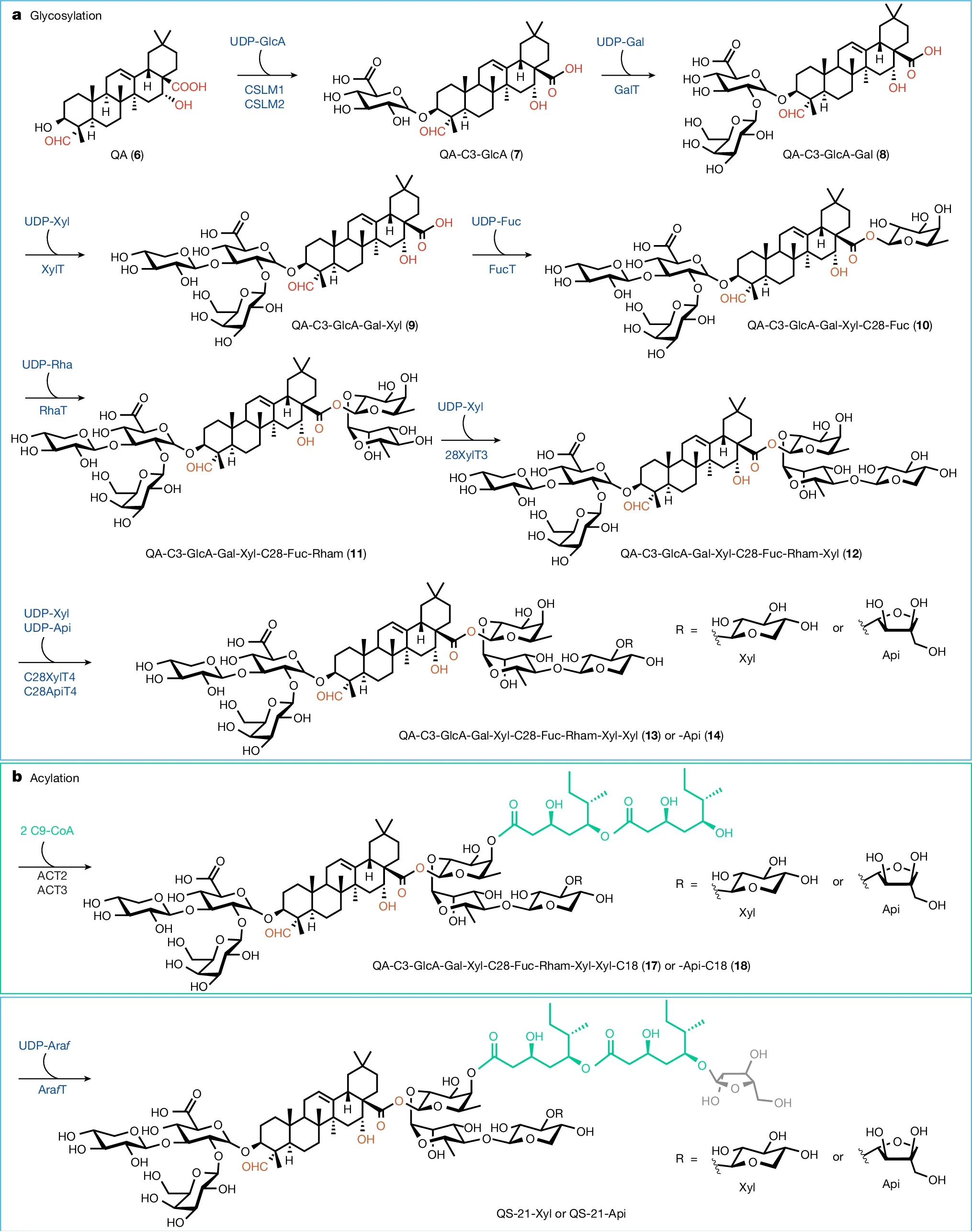
Figure: Complete biosynthetic pathway for the de novo production of QS-21 in yeast from simple sugars (Liu et al., 2024).
Yes, baker’s yeast could produce QS-21, but the production levels were orders of magnitude below commercial viability. This is the challenge that Maria and her team set out to solve.
Why does yeast struggle to produce QS-21, despite using the exact same enzymes as those used by the soap bark tree? It’s not surprising if you consider the evolutionary pressures that shaped these enzymes. They evolved to function optimally in the cells of the soap bark tree, growing in the cool wet winters and hot dry summers of the Chilean forest. QS-21 pathway enzymes operate in different compartments of the plant cell, each with its own metabolite concentrations and pH. When transferred into a yeast cell and exposed to the temperatures, pH and pressures of an industrial fermentation vessel, these enzymes falter.
Activity values are low. Key enzymes show poor processivity, too often releasing partially finished reaction intermediates rather than neatly completing the reaction. Misfolding could occur, or poor membrane insertion. And enzymes could struggle to remain stable and functional with the pH shifts, temperature fluctuations, or oxidative stress of an industrial process. The solution lies in enzyme engineering, to improve activity, processivity and resolve the main pathway bottleneck. The problem is: evolution had many thousands of years to iterate on these enzymes. With a growing vaccine demand, Quillaja saponaria ain’t got time for that.
Why machine learning fits this optimization problem
Traditional directed evolution would require screening tens of thousands of variants per enzyme, multiplied across a multi-enzyme pathway. The numbers become prohibitive quickly. Rational design faces different constraints - scientists only get so far when trying to predict how sequence changes affect processivity and product specificity for large, multidomain enzymes.
The Keasling Lab integrated Cradle into their optimization workflow to address these constraints. Machine learning is useful here because it learns sequence-function relationships directly from experimental data without requiring mechanistic understanding upfront. The models propose variants based on patterns in how sequence changes affect measured properties (catalytic activity, product specificity, thermal stability under fermentation conditions), and each experimental round improves the predictions.
These enzymes take decades to make enough of QS-21. We need to optimize their speed and their activity, their processivity. There are so many features that need to be optimized in order to be able to produce QS-21 at scale.
Maria Astolfi
The workflow runs in iterative cycles. Cradle generates diverse sequence libraries predicted to improve target properties, the Keasling Lab expresses and characterizes them, and the experimental results feed back to retrain the models. Cradle's platform handles sequence generation and property prediction in a unified workflow, which matters when optimizing multiple enzymes in parallel while maintaining pathway balance.
Early results and remaining bottlenecks
The team is seeing measurable improvements in individual enzyme properties. Initial rounds have identified variants with strong improvements in catalytic efficiency processivity under fermentation conditions. But the real test is whether they can coordinate improvements across the entire pathway to achieve commercially viable titers while maintaining product fidelity.
That last point is critical. QS-21 has a precise molecular structure, including its glycosylation pattern and the linkages connecting its domains, that determines its immunological properties. Any structural deviation affects adjuvant activity. The researchers need to optimize for higher titers of the exact molecule.
The downstream challenges remain significant. Even with optimized enzymes, the team will need to solve purification (QS-21 is structurally similar to dozens of other saponins produced by the same pathway), demonstrate batch-to-batch consistency at a level acceptable for pharmaceutical manufacturing, and navigate regulatory approval for a biosynthetic version of a molecule currently extracted from plants.
But the traditional alternative isn't viable long-term. Vaccine demand is growing. The population of harvestable Quillaja saponaria trees in Chile is not. And extraction-based supply chains are vulnerable to climate variability, geopolitical instability, and biodiversity loss.
A systematic approach to plant enzyme optimization
The broader question is whether researchers can develop a systematic approach to optimizing plant-derived enzymes for industrial microbial production. Hundreds of high-value molecules are currently extracted from plants, many from biodiversity hotspots where harvesting pressure threatens ecosystems. Synthetic biology offers sustainable alternatives, but plant enzymes consistently underperform when moved into heterologous hosts.
If machine learning can accelerate the optimization process—reducing screening requirements from tens of thousands of variants to hundreds—it changes the economics of replacing extraction with fermentation. The cost structure shifts from marginal improvements on an inherently unsustainable process to establishing new production routes that scale with fermentation capacity.
That's the bet the Keasling Lab is making with QS-21. The early data suggests it's worth pursuing, and the methodology should transfer to other plant-derived pharmaceuticals facing similar supply chain constraints. The Keasling Lab is currently working on the second round of generated sequences after which it plans to run a third. Intermediary data and results will be published soon in their preprint on bioRxiv.
Maria Astolfi is a synthetic biologist in the Keasling Lab at UC Berkeley, where her work focuses on sustainable biomanufacturing of pharmaceuticals currently extracted from plants.
Image rights: Quillaja saponaria 09.jpg by Yastay is licensed under CC BY-SA 4.0
The “soap bark tree” Quillaja saponaria gets its name from the saponins in its bark—natural compounds that create a soapy lather in water. Indigenous people in the Andes have used this bark for generations as a natural remedy. Scientists later discovered that one of these saponins, a molecule called QS-21, could act as a ‘vaccine adjuvant’ to boost the immune system's response to vaccines. Today, QS-21 is a key ingredient in vaccines against shingles, malaria, and RSV.
As vaccine demand grows, so does pressure on Chile's soap bark tree populations. The trees are already threatened by deforestation and wildfires, and they grow slowly—taking decades to reach maturity. While sustainable harvesting methods have been introduced, meeting pharmaceutical demand at scale remains difficult. Biologists at UC Berkeley are working on a solution.

Maria Astolfi, born and raised in the Amazonian rainforest, is a synthetic biologist in the Keasling Lab at UC Berkeley. She’s part of a research project to replace this extraction-based supply chain with precision fermentation, and uses Cradle to accelerate their research.
Plant enzymes in a yeast cell
In precision fermentation, researchers engineer a microbe like baker’s yeast so that it becomes able to ‘brew’ a specific molecule of interest in a fermentation vat. In 2024, researchers from the Keasling lab demonstrated that it was possible to produce QS-21 using this approach (published in Nature in 2024, source). They added a copy of 38 plant genes from Quillaja saponaria into baker’s yeast, engineering its metabolism to now be able to convert renewable resources like simple sugars into this sought-after vaccine adjuvant.

Figure: Complete biosynthetic pathway for the de novo production of QS-21 in yeast from simple sugars (Liu et al., 2024).
Yes, baker’s yeast could produce QS-21, but the production levels were orders of magnitude below commercial viability. This is the challenge that Maria and her team set out to solve.
Why does yeast struggle to produce QS-21, despite using the exact same enzymes as those used by the soap bark tree? It’s not surprising if you consider the evolutionary pressures that shaped these enzymes. They evolved to function optimally in the cells of the soap bark tree, growing in the cool wet winters and hot dry summers of the Chilean forest. QS-21 pathway enzymes operate in different compartments of the plant cell, each with its own metabolite concentrations and pH. When transferred into a yeast cell and exposed to the temperatures, pH and pressures of an industrial fermentation vessel, these enzymes falter.
Activity values are low. Key enzymes show poor processivity, too often releasing partially finished reaction intermediates rather than neatly completing the reaction. Misfolding could occur, or poor membrane insertion. And enzymes could struggle to remain stable and functional with the pH shifts, temperature fluctuations, or oxidative stress of an industrial process. The solution lies in enzyme engineering, to improve activity, processivity and resolve the main pathway bottleneck. The problem is: evolution had many thousands of years to iterate on these enzymes. With a growing vaccine demand, Quillaja saponaria ain’t got time for that.
Why machine learning fits this optimization problem
Traditional directed evolution would require screening tens of thousands of variants per enzyme, multiplied across a multi-enzyme pathway. The numbers become prohibitive quickly. Rational design faces different constraints - scientists only get so far when trying to predict how sequence changes affect processivity and product specificity for large, multidomain enzymes.
The Keasling Lab integrated Cradle into their optimization workflow to address these constraints. Machine learning is useful here because it learns sequence-function relationships directly from experimental data without requiring mechanistic understanding upfront. The models propose variants based on patterns in how sequence changes affect measured properties (catalytic activity, product specificity, thermal stability under fermentation conditions), and each experimental round improves the predictions.
These enzymes take decades to make enough of QS-21. We need to optimize their speed and their activity, their processivity. There are so many features that need to be optimized in order to be able to produce QS-21 at scale.
Maria Astolfi
The workflow runs in iterative cycles. Cradle generates diverse sequence libraries predicted to improve target properties, the Keasling Lab expresses and characterizes them, and the experimental results feed back to retrain the models. Cradle's platform handles sequence generation and property prediction in a unified workflow, which matters when optimizing multiple enzymes in parallel while maintaining pathway balance.
Early results and remaining bottlenecks
The team is seeing measurable improvements in individual enzyme properties. Initial rounds have identified variants with strong improvements in catalytic efficiency processivity under fermentation conditions. But the real test is whether they can coordinate improvements across the entire pathway to achieve commercially viable titers while maintaining product fidelity.
That last point is critical. QS-21 has a precise molecular structure, including its glycosylation pattern and the linkages connecting its domains, that determines its immunological properties. Any structural deviation affects adjuvant activity. The researchers need to optimize for higher titers of the exact molecule.
The downstream challenges remain significant. Even with optimized enzymes, the team will need to solve purification (QS-21 is structurally similar to dozens of other saponins produced by the same pathway), demonstrate batch-to-batch consistency at a level acceptable for pharmaceutical manufacturing, and navigate regulatory approval for a biosynthetic version of a molecule currently extracted from plants.
But the traditional alternative isn't viable long-term. Vaccine demand is growing. The population of harvestable Quillaja saponaria trees in Chile is not. And extraction-based supply chains are vulnerable to climate variability, geopolitical instability, and biodiversity loss.
A systematic approach to plant enzyme optimization
The broader question is whether researchers can develop a systematic approach to optimizing plant-derived enzymes for industrial microbial production. Hundreds of high-value molecules are currently extracted from plants, many from biodiversity hotspots where harvesting pressure threatens ecosystems. Synthetic biology offers sustainable alternatives, but plant enzymes consistently underperform when moved into heterologous hosts.
If machine learning can accelerate the optimization process—reducing screening requirements from tens of thousands of variants to hundreds—it changes the economics of replacing extraction with fermentation. The cost structure shifts from marginal improvements on an inherently unsustainable process to establishing new production routes that scale with fermentation capacity.
That's the bet the Keasling Lab is making with QS-21. The early data suggests it's worth pursuing, and the methodology should transfer to other plant-derived pharmaceuticals facing similar supply chain constraints. The Keasling Lab is currently working on the second round of generated sequences after which it plans to run a third. Intermediary data and results will be published soon in their preprint on bioRxiv.
Maria Astolfi is a synthetic biologist in the Keasling Lab at UC Berkeley, where her work focuses on sustainable biomanufacturing of pharmaceuticals currently extracted from plants.
Image rights: Quillaja saponaria 09.jpg by Yastay is licensed under CC BY-SA 4.0
Subscribe to new case studies and other Cradle updates
Recent posts
Subscribe and get new posts and updates from Cradle straight to your inbox.
Follow Cradle
Built with ❤️ in Amsterdam & Zurich
Follow Cradle
Built with ❤️ in Amsterdam & Zurich
Follow Cradle