Engineering CRISPR on-target editing activity from <25% to 68% in three rounds
Scientists using Cradle optimized a CRISPR system for on-target editing activity across three rounds. After a prior screening campaign of >3,000 variants achieved best-in-class performance of <25%, CRADLE-1 delivered candidates reaching 50%, 56%, and 68% on-target activity in successive rounds.
Modality | CRISPR gene-editing system |
Target | Redacted |
Properties optimized | On-target editing activity |
Rounds | 3 |
Candidates per round | 96 |
Key result | 68% on-target editing (vs <25% from prior screening) |
Partner | Commercial partner |
Data availability | Redacted |
Context
CRISPR-based gene editing systems have revolutionized molecular biology and hold therapeutic potential for genetic diseases. Editing efficiency—the percentage of target cells successfully modified—directly determines the feasibility of therapeutic applications. Many CRISPR systems exhibit modest on-target activity in primary cells or challenging genomic contexts, limiting their clinical utility.
Improving on-target activity without generating off-target edits or cytotoxicity represents a critical engineering challenge for translating CRISPR therapeutics from research tools to approved medicines.
Challenge
The commercial partner had previously run an extensive screening campaign testing >3,000 variants of the CRISPR system. Despite this large-scale effort, the best variant achieved on-target editing activity of <25%—insufficient for the therapeutic application. The partner provided this complete dataset to Cradle, representing substantial investment in sequence-function measurements that had not yet yielded a clinically viable candidate.
The challenge was to extract signal from this dataset—which demonstrated that the vast majority of sequence space yielded poor editors—and identify the rare beneficial mutations that could push activity into the therapeutically relevant regime (>50%).
Approach
We performed three rounds of Cradle optimization with 96 candidates each. All rounds consumed the full >3,000-variant screening dataset as training data. Round 1 was configured with on-target editing activity as the primary objective, using supervised predictors trained on the screening results to identify promising regions of sequence space.
The supervised predictors were trained on the partner's historical data, where the best observed on-target editing activity was <25%. Despite this modest starting point, Cradle was able to learn from the available sequence-function relationships and generate variants that substantially exceeded previous results — reaching 50% in the first round and ultimately 68% by the third.
Results
Cradle achieved progressive improvements across three rounds, successfully extrapolating beyond the screening campaign's best result:
Round | Best on-target editing activity | Fold improvement over screening |
Screening (>3,000 variants) | <25% | Baseline |
Cradle Round 1 | 50% | 2.0× |
Cradle Round 2 | 56% | 2.2× |
Cradle Round 3 | 68% | 2.7× |
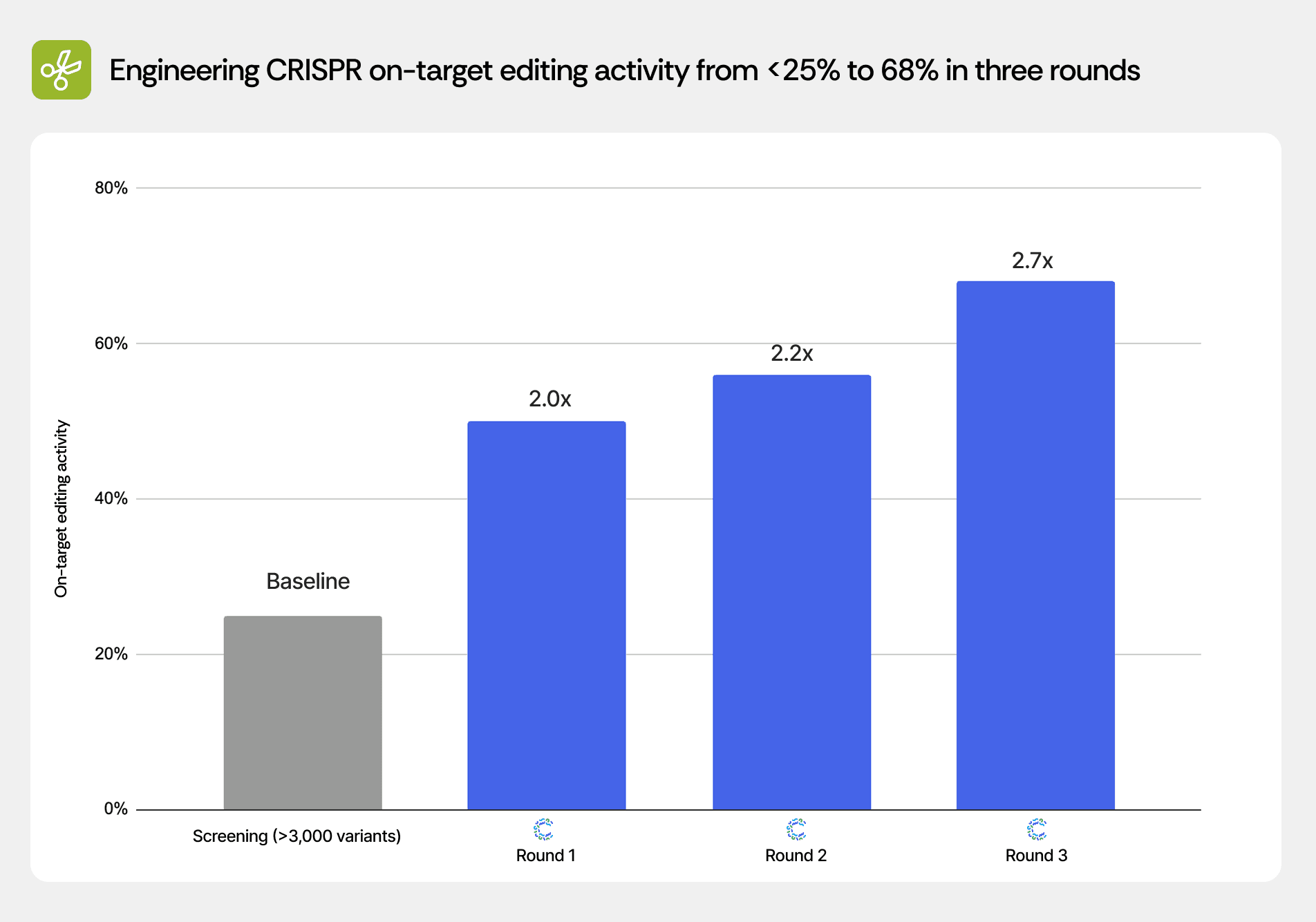
The Round 1 result of 50% activity doubled the screening campaign's best performance, demonstrating that Cradle successfully identified beneficial mutations not sampled in the >3,000-variant library. Rounds 2 and 3 continued to push activity higher, suggesting that the initial >3,000 variants had not exhausted the optimization landscape and that systematic machine learning guidance could navigate toward higher-performing regions.
The progressive round-over-round improvements (50% → 56% → 68%) indicate that Cradle's learning from wet lab data in Rounds 1 and 2 enabled increasingly accurate predictions in Rounds 2 and 3. This contrasts with the screening campaign, which tested a large number of variants without systematic feedback-driven refinement.
The final 68% activity represents a clinically meaningful threshold for many therapeutic applications, transforming an editing system with marginal utility into one approaching practical deployment.
What this means
This result demonstrates Cradle's ability to learn from and improve upon large-scale screening efforts. Rather than treating the >3,000-variant dataset as a failed campaign, Cradle extracted sufficient signal to predict substantially better variants. The 2.7-fold improvement achieved with 288 total candidates compares favorably to the >3,000 variants tested via screening, representing both cost efficiency and superior outcomes.
The ability to extrapolate beyond the training distribution (predicting 68% activity when trained on data capped at <25%) suggests that the supervised predictors learned generalizable sequence-function relationships rather than simply memorizing the training examples. This capability is critical for lead optimization, where the goal is always to exceed the best available candidate.
For gene therapy development, achieving on-target editing rates above 50–60% can be the difference between a research tool and a therapeutic candidate, making this improvement commercially and clinically significant.
Methods note
Cradle ingested the complete >3,000-variant screening dataset for supervised predictor training. Evotuning was performed on the evolutionary alignment of the CRISPR system. On-target editing activity was measured via the partner's proprietary cellular assay. Generation used diversity-aware ranking to balance exploitation of predicted high-activity regions with exploration of undersampled sequence space. Full details of the CRISPR system and editing assay remain confidential to the commercial partner.
Recent posts
Subscribe and get new posts and updates from Cradle straight to your inbox.