Engineering polyspecific anti-venom VHHs with sub-nanomolar binding and high thermostability
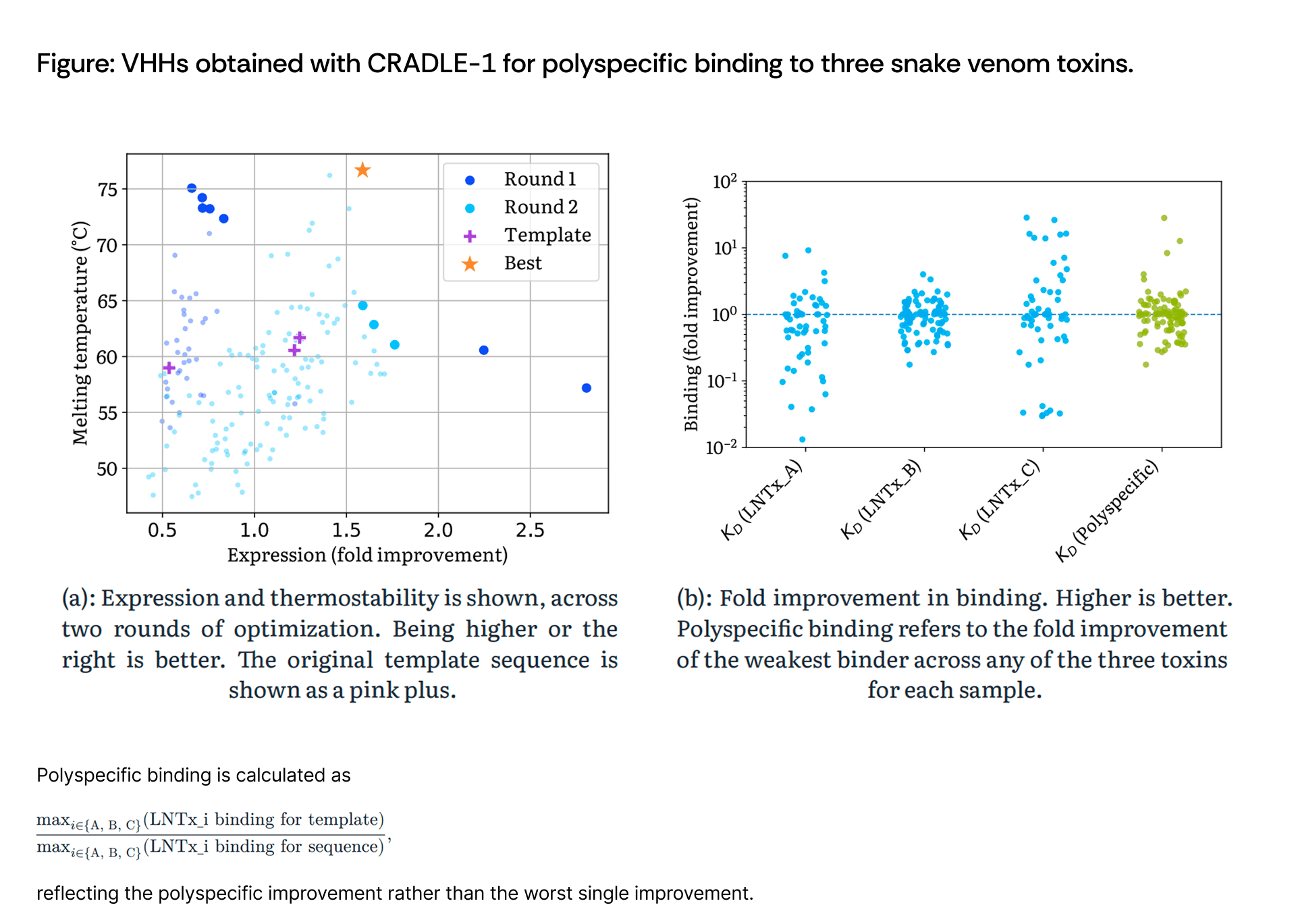
Scientists using Cradle optimized VHHs for polyspecific binding to three distinct snake venom neurotoxins, thermostability, and expression across two rounds. The best candidate achieved binding below 100 pM, below 1 nM, and below 2 nM to the three toxins respectively, with a melting temperature of 76.7 °C and 1.59× improved expression.
Modality | VHH (nanobody) |
Target | Three long-chain three-finger α-neurotoxins (LNTx A, B, C) |
Properties optimized | Binding (3×), thermostability, expression |
Rounds | 2 |
Candidates per round | 96 |
Key result | K_D <100 pM (LNTx A), <1 nM (LNTx B), <2 nM (LNTx C), T_m 76.7 °C |
Partner | Academic collaboration |
Data availability | Available in upcoming publication |
Context
Snakebites cause an estimated 58,000 deaths annually in India and 7,000 in sub-Saharan Africa. The species of snake responsible for a bite may not always be identified when administering treatment, making polyvalent antivenoms critical for effective therapy. Additionally, cold storage infrastructure is often unavailable in regions where snakebites are most common, creating a strong need for thermostable biologics that can withstand ambient temperatures.
Long-chain three-finger α-neurotoxins (LNTxs) are common among elapid snakes, making them high-value targets for broad-spectrum antivenom development.
Challenge
Three template sequences were initially obtained using Cradle’s hit identification pipeline applied to phage and yeast display data. The optimization required simultaneous binding to three structurally related but distinct neurotoxins—a five-property optimization problem when including thermostability and expression. This represents a particularly challenging multi-objective landscape with potential trade-offs between binding to different toxins.
The requirement for high thermostability (to enable distribution without cold chain infrastructure) further constrained the viable sequence space, as stability-enhancing mutations can often reduce binding affinity or expression.
Approach
Scientists performed two rounds of Cradle optimization with 96 candidates each, making mutations to both CDR and framework regions. Round 1 operated as a zero-shot diversification starting from the three template sequences. Round 2 optimized thermostability as the primary objective while setting binding and expression as relative constraints (tasked to improve relative to the templates).
The optimization leveraged screening data from the phage and yeast display campaigns as in-domain context. While this data provided sequences without quantitative binding values, it enriched the evolutionary context with CDR-region diversity, which is particularly valuable for antibody optimization where CDRs are not evolutionarily conserved.
Results
The final candidate achieved exceptional polyspecific binding alongside high thermostability and improved expression:
Property | Best Cradle candidate |
K_D (LNTx A) | <100 pM |
K_D (LNTx B) | <1 nM |
K_D (LNTx C) | <2 nM |
T_m | 76.7 °C |
Expression | 1.59× relative to templates |
The polyspecific binding profile demonstrates broad neutralization potential across structurally related neurotoxins, a critical requirement for polyvalent antivenoms where the specific toxin composition varies by snake species and geographic region. The melting temperature of 76.7 °C substantially exceeds typical antibody stability, enabling potential formulation strategies for distribution without cold chain requirements.
Expression improvement of 1.59× over the already-functional templates ensures manufacturability at scale, an important consideration for therapies targeting neglected tropical diseases where cost-effectiveness is paramount.
What this means
This work demonstrates Cradle's effectiveness for optimizing therapeutic antibodies targeting neglected tropical diseases. The combination of polyspecific sub-nanomolar binding with exceptional thermostability addresses two critical challenges for antivenom development: broad coverage across toxin variants and stability in resource-limited settings.
The ability to optimize five properties simultaneously (three binding targets plus thermostability and expression) from screening data, achieving clinically relevant results in just two rounds, suggests that Cradle can accelerate development timelines for global health applications where resources for extensive optimization campaigns are limited.
Methods note
Cradle used evotuning on both evolutionary alignments and screening sequences from the phage/yeast display campaigns. Enrichment fold-changes from screening were used to supervise binding affinity predictions. Both CDR and framework mutations were permitted. Binding kinetics were measured via SPR, thermostability via DSF, and expression in E. coli quantified via LabChip protein characterization. The academic partner will release full experimental details in an upcoming publication. Methodological details are available in the CRADLE-1 whitepaper.
Recent posts
Subscribe and get new posts and updates from Cradle straight to your inbox.